
Bacterial and Viral Bioinformatic Resource Center Emerges as Global Hub for Antimicrobial Resistance Training
Infectious pathogens in bacteria, viruses, fungi, and parasites naturally evolve over time. As they do, they can develop antimicrobial resistance (AMR) to available medicines. When this happens, antibiotics and other antimicrobial medicines lose efficacy and these infections become more difficult to treat, increasing the likelihood of disease spread, severity, and death.
Drug-resistant pathogens can be found all around us – in people, animals, food, and the environment. Their emergence and rapid spread are an ever-present and growing global concern. According to the World Health Organization (WHO), resistance against antibiotics frequently used to treat common infections like urinary tract infections, sepsis, and diarrhea, is occurring worldwide. Further exacerbating the problem is the emergence of "superbugs" – multi- and pan-resistant infectious bacteria that are unresponsive to any existing antimicrobial medications.
As the threat worsens, the UVA Biocomplexity Institute and its team are responding through their work on the Bacterial and Viral Bioinformatics Resource Center (BV-BRC). The BV-BRC is an integrated resource of computational systems that collect, compare, and analyze genomic-level data relevant to infectious diseases, pathogens, and their interactions. It was launched by the National Institute of Allergy and Infectious Diseases (NIAID) in 2004, providing researchers worldwide with free access to sophisticated tools to address important health issues, ranging from AMR and perennial diseases to emerging epidemics and bioengineered threats.
Originally started as part of the Department of Homeland Security's biodefense program, the BV-BRC has evolved significantly over the last 19 years. What began as a modest platform where researchers could simply view data related to eight bacterial genomes has expanded into a comprehensive database and end-to-end suite of analysis tools – a one-stop shop – to study more than 723,000 bacterial genomes and over 9.5 million viral genomes.
The BV-BRC combines the data, computational tools, and technologies from three legacy centers used by the scientific community: PATRIC, which supplied bacterial data, and IRD and ViPR, which supplied viral data. In 2019, the National Institutes of Health awarded the Biocomplexity Institute and its partners a contract to consolidate the three centers into one.
Today, the BV-BRC has more than 40,000 registered users around the world. These users include students, biological and clinical researchers, epidemiologists, data scientists, and professionals from academia, pharmaceutical companies, food and beverage, public health, government labs, and agencies such as the Centers for Disease Control and Prevention (CDC), U.S. Food and Drug Administration (FDA), and U.S. Department of Agriculture (USDA).
"Among many other types of analyses, the BV-BRC enables researchers all over the world to take any metagenomic sample – from waste water, the clinical setting, and so on – and better understand, from a biological standpoint, what agents are present," said Rebecca Wattam, Research Associate Professor for the Biocomplexity Institute's Network Systems Science and Advanced Computing (NSSAC) division. "The resource provides tools to pull out complete genomes, compare them to hundreds of thousands of other known bacteria and viral organisms, and flag critical characteristics such as antimicrobial resistance. Once researchers understand more about the genomics of certain bacteria or viruses, more effective antimicrobial medications or vaccines can be developed to help treat those infections."
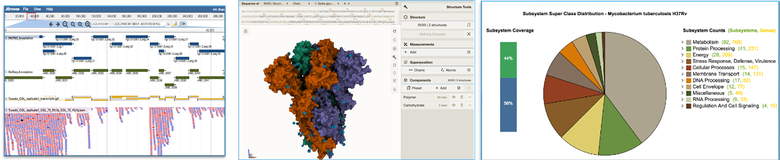 A sampling of the visual analytics tools included in the BV-BRC. The free, open-access data and tools are available to researchers worldwide.
A sampling of the visual analytics tools included in the BV-BRC. The free, open-access data and tools are available to researchers worldwide.Today, the Biocomplexity Institute is a global leader in antimicrobial resistance workshops and training, and has hosted thousands of attendees at nearly 100 workshops over the last two decades in Charlottesville and locations throughout the United States, Asia, Europe, Africa, and South America. This year, the Institute hosted more than 20 attendees for a two-day workshop in April and is planning additional dates this fall. Through these events, the Institute team trains the world’s researchers to use the BV-BRC to address the AMR-related questions and issues that are most pressing to their work and respective needs.
"People are looking for answers, not just a tutorial of the tools," said Wattam, the lead for BV-BRC outreach and training. "In our workshops, we ask attendees what questions they have, then show them how to use the BV-BRC resource to find their answers."
"Recently, we had data from a mixed sample of a bacterial infection resulting from a hip replacement. We can take the samples and identify the pathogens present in the sample and isolate their genomes. We conduct DNA- and protein-level analysis on these genomes so that people can compare them to others available in the resource and determine what is different and how that may affect people, cause illness, and potentially identify drugs or treatments to slow them down. We focus on real-world use cases, so researchers can walk away with a practical knowledge of how to use the BV-BRC to find the answers, and potentially, the solutions they need."
Staying on the forefront of innovation, the BV-BRC integrates machine learning and artificial intelligence techniques to quickly analyze data and predict AMR and other red-flag threats. Based on the hundreds of thousands of bacterial genomes that have been sequenced and analyzed, the BV-BRC can take a particular bacterial genome sequence and predict how it will respond to certain antibiotics. Its ability to quickly and thoroughly analyze the large spectrum of data related to pathogens makes the BV-BRC a standout resource for infectious disease researchers.
"The BV-BRC is truly unique in that it brings comprehensive data and analysis together in one user-friendly platform," said Ron Kenyon, senior scientist in the Institute's NSSAC division. "It now also has the ability to analyze genomic sequence read quality to ensure that the information provided about a particular genome is accurate. Previously, researchers may not have been aware of inaccuracies until they were weeks into the process of analyzing. Now, we have instant warning flags along the way."
The National Institutes of Health (NIH) and National Institute of Allergy and Infectious Diseases (NIAID) has consistently funded the BV-BRC since 2004. The resource is funded primarily for bacterial and pathogen researchers, but can also be used for dedicated projects addressing chronic or emerging diseases and epidemics such as COVID, MERS, and Ebola.