Topological Complexity of Interacting Systems
The complexity of an interacting system also arises from its "shape", i.e., its topological complexity. Cross-serial interactions (pseudoknots), as well as multiple-structure interactions (riboswitches), are crucial to the function of biomolecules. The increased complexity of these interactions brings new challenges as well as hints at new methods for understanding them. We construct mathematical models to measure the topological complexity of interacting systems and tackle such complexity using topological recursion. Our research can facilitate the detection and design of functional genes whose structures rank high on this topological complexity scale.
Research Highlights
- We apply simplicial homology and study the topological trace between two RNA secondary structures. Such a trace captures the mutually exclusive substructures that enable the "switching" mechanism in riboswitches. This also can be used to distinguish ncRNAs from different classes.
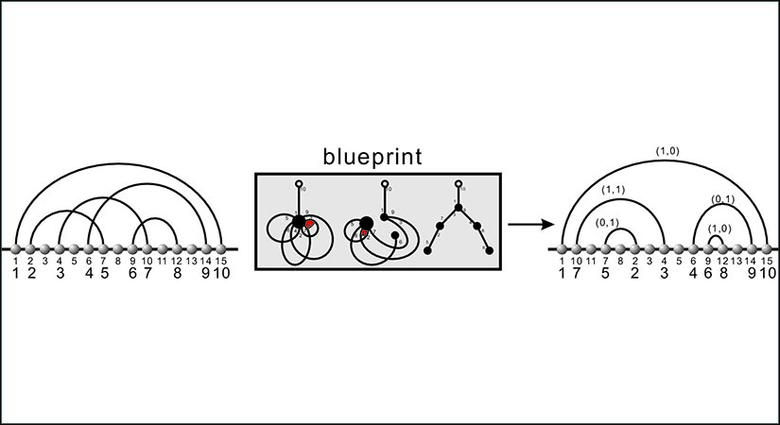
- We derived a novel method for transforming cross-serial interactions into cross-free interactions. The method facilitates fast Boltzmann sampling and statistical analysis of RNA pseudoknot structures.
Ongoing Research
- We are developing topological approaches for identifying mutually exclusive substructures in the structure ensemble of a given sequence, locating switching sequences, and detecting potential riboswitches.
- We are designing efficient algorithms to construct riboswitch sequences with two desired stable configurations.
- We are extending the homology analysis to planar interaction structures in order to understand the interplay between ligand binding and folding.
Publications
Mathematical Biocomplexity
Mathematical Biocomplexity
Article
Topological language for RNA
Mathematical Biocomplexity
Mathematical Biocomplexity