Network Systems Science and Advanced Computing

Our Team
Our team integrates diverse research perspectives. We're cognitive, behavioral and social scientists, along with epidemiologists, data analysts, statisticians, biochemists and software engineers. Working collaboratively, we pool our capabilities and theoretical knowledge to tackle stubborn and evolving issues in the world, from contagion of behaviors and diseases to disruptions in resource distribution and ecosystems.
Our Areas of Focus
We apply our computational modeling tools and combined expertise to tackle a broad swath of issues affecting human life – from improving disaster resiliency in city infrastructures to identifying previously unknown organisms in our environments and bodies.
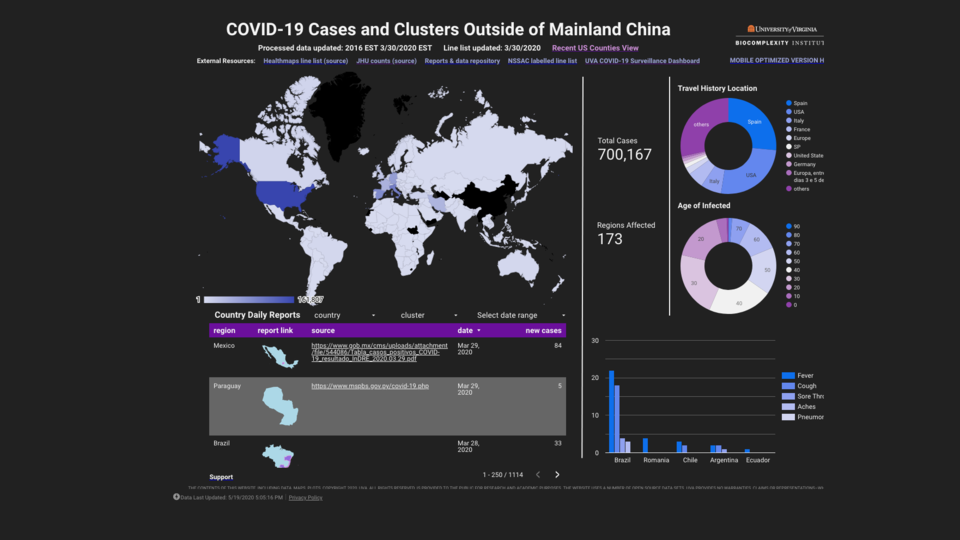
Modeling for COVID-19
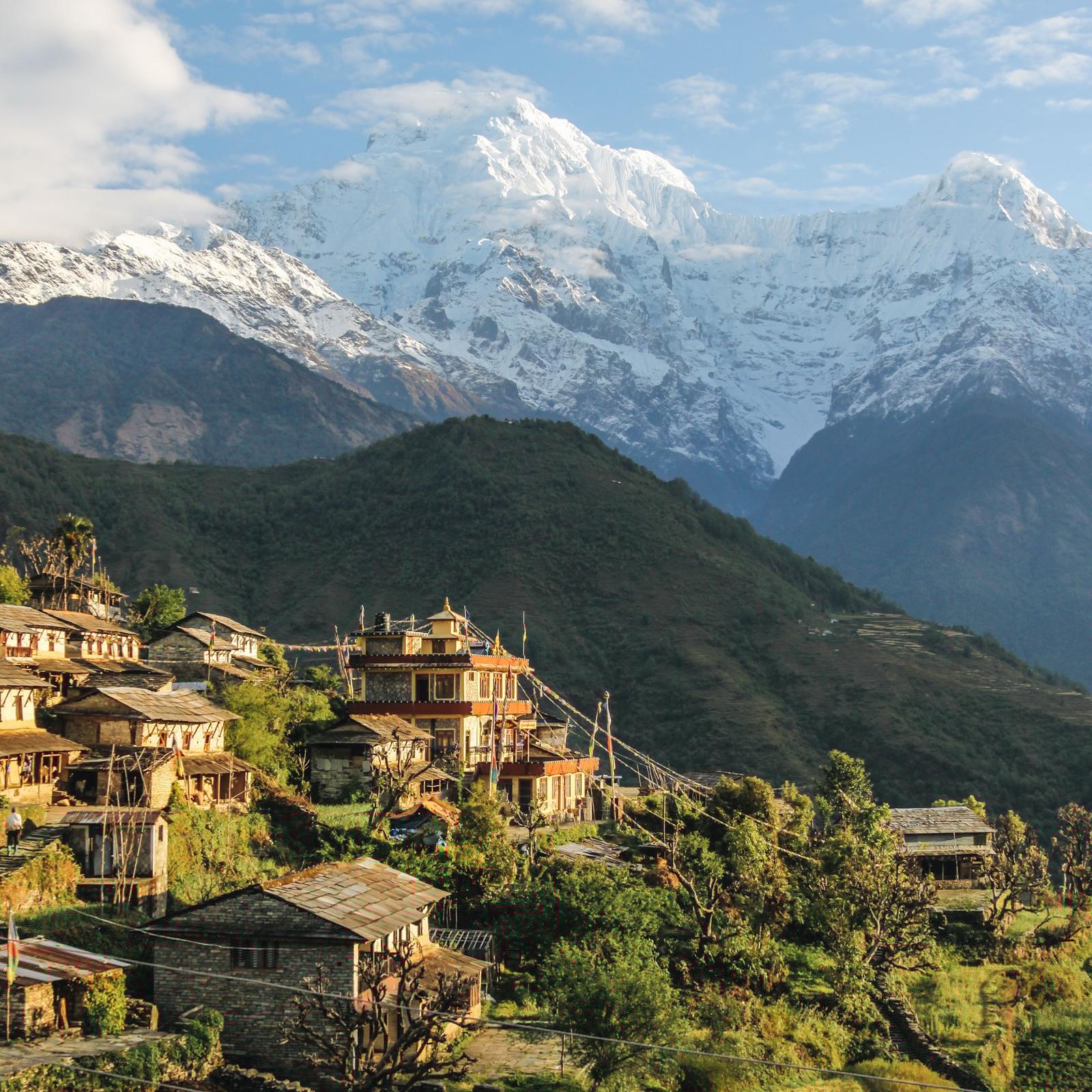
We've modeled several epidemics in the past 20 years, including H1N1 and Ebola. Today, collaborating with researchers from across the Institute and the University of Virginia, we're modeling scenarios to help governments and the global research community understand and mitigate the spread of COVID-19.
Computing for Global Challenges Program
From human health to infrastructure, our world is facing enormous threats. To understand them, we need and use data. This undergraduate program is a training ground for the next generation of data-driven researchers. Working in teams with peers and scientists from the Institute's Network Systems Science and Advanced Computing division, students learn a new way to address real-world issues while working on critical research projects.

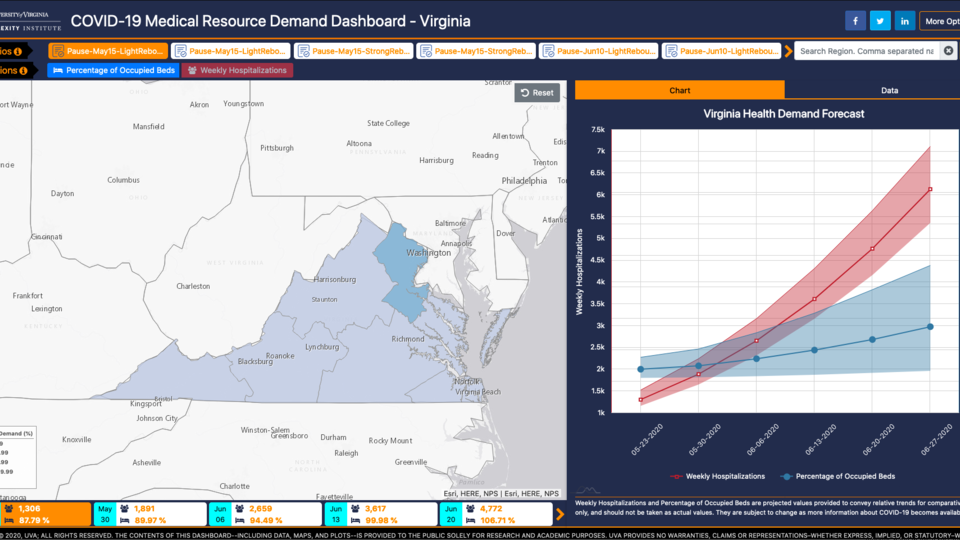
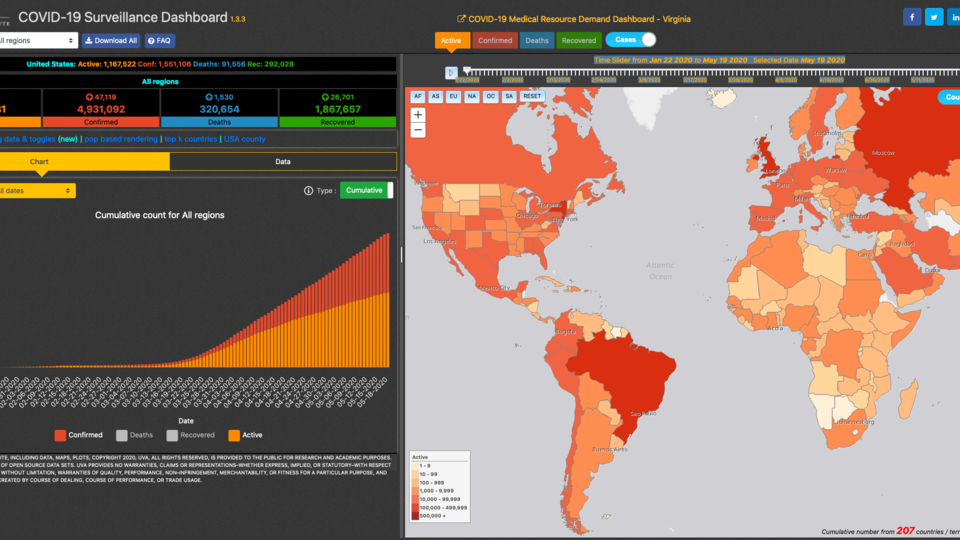
Use Our Tools
We develop tools such as dashboards and data sets that we make available to researchers, academics, policymakers and the public.
Work with Us
We are able to do the type of research we do because of the people we partner with, and the people we hire.
We’re fortunate to work with global partners and sponsors from all segments of government, global foundations, academia, and industry to solve society’s most complex challenges. We approach these problems by assembling teams of experts in a variety of fields to work together and create solutions that challenge the very fields in which our teams operate.